YamboConvergence: fixed-path¶
The YamboConvergence workchain provides the functionalities to run multiple G0W0 calculations on the same system, over a wide range of chaging parameter.
This represents the typical method to obtain an accurate evaluation of the quasiparticle correction: indeed, a lot of effort has to be done in order to find
the convergence with respect to parameters like empty states used to evaluate the Self Energy used to solve the quasiparticle equation.
There are cases in which convergences cannot be achieved exactly: this is, e.g., the case of molecules. In such a case, you may want to perform G0W0
calculations on a give set of parameters/values and then, with an extrapolation, obtain the given result.
This logic can be activated in the YamboConvergence, as we will see, and represents a new implementation that for now has not the extrapolation feature
included.
Let’s see the case of fixed-path automation, activated by setting "type": 2D_space:
#!/usr/bin/env runaiida
# -*- coding: utf-8 -*-
from __future__ import absolute_import
from __future__ import print_function
import sys
import os
from aiida.plugins import DataFactory, CalculationFactory
from aiida.orm import List, Dict
from aiida.engine import submit
from aiida_yambo.workflows.yamboconvergence import YamboConvergence
from aiida_quantumespresso.utils.pseudopotential import validate_and_prepare_pseudos_inputs
from ase import Atoms
import argparse
def get_options():
parser = argparse.ArgumentParser(description='YAMBO calculation.')
parser.add_argument(
'--yambocode',
type=int,
dest='yambocode_pk',
required=True,
help='The yambo(main code) codename to use')
parser.add_argument(
'--parent',
type=int,
dest='parent_pk',
required=False,
help='The parent to use')
parser.add_argument(
'--yamboprecode',
type=int,
dest='yamboprecode_pk',
required=True,
help='The precode to use')
parser.add_argument(
'--pwcode',
type=int,
dest='pwcode_pk',
required=True,
help='The pw to use')
parser.add_argument(
'--pseudo',
type=str,
dest='pseudo_family',
required=True,
help='The pseudo_family')
parser.add_argument(
'--time',
type=int,
dest='max_wallclock_seconds',
required=False,
default=30*60,
help='max wallclock in seconds')
parser.add_argument(
'--nodes',
type=int,
dest='num_machines',
required=False,
default=1,
help='number of machines')
parser.add_argument(
'--mpi',
type=int,
dest='num_mpiprocs_per_machine',
required=False,
default=1,
help='number of mpi processes per machine')
parser.add_argument(
'--threads',
type=int,
dest='num_cores_per_mpiproc',
required=False,
default=1,
help='number of threads per mpi process')
parser.add_argument(
'--queue_name',
type=str,
dest='queue_name',
required=False,
default=None,
help='queue(PBS) or partition(SLURM) name')
parser.add_argument(
'--qos',
type=str,
dest='qos',
required=False,
default=None,
help='qos name')
parser.add_argument(
'--account',
type=str,
dest='account',
required=False,
default=None,
help='account name')
args = parser.parse_args()
###### setting the machine options ######
options = {
'yambocode_pk': args.yambocode_pk,
'yamboprecode_pk': args.yamboprecode_pk,
'pwcode_pk': args.pwcode_pk,
'pseudo_family': args.pseudo_family,
'max_wallclock_seconds': args.max_wallclock_seconds,
'resources': {
"num_machines": args.num_machines,
"num_mpiprocs_per_machine": args.num_mpiprocs_per_machine,
"num_cores_per_mpiproc": args.num_cores_per_mpiproc,
},
'custom_scheduler_commands': u"export OMP_NUM_THREADS="+str(args.num_cores_per_mpiproc),
}
if args.parent_pk:
options['parent_pk']=args.parent_pk
if args.queue_name:
options['queue_name']=args.queue_name
if args.qos:
options['qos']=args.qos
if args.account:
options['account']=args.account
return options
def main(options):
###### setting the lattice structure ######
alat = 2.4955987320 # Angstrom
the_cell = [[1.000000*alat, 0.000000, 0.000000],
[-0.500000*alat, 0.866025*alat, 0.000000],
[0.000000, 0.000000, 6.4436359260]]
atoms = Atoms('BNNB', [(1.2477994910, 0.7204172280, 0.0000000000),
(-0.0000001250, 1.4408346720, 0.0000000000),
(1.2477994910, 0.7204172280, 3.2218179630),
(-0.0000001250,1.4408346720, 3.2218179630)],
cell = [1,1,1])
atoms.set_cell(the_cell, scale_atoms=False)
atoms.set_pbc([True,True,True])
StructureData = DataFactory('structure')
structure = StructureData(ase=atoms)
###### setting the kpoints mesh ######
KpointsData = DataFactory('array.kpoints')
kpoints = KpointsData()
kpoints.set_kpoints_mesh([6,6,2])
###### setting the scf parameters ######
Dict = DataFactory('dict')
params_scf = {
'CONTROL': {
'calculation': 'scf',
'verbosity': 'high',
'wf_collect': True
},
'SYSTEM': {
'ecutwfc': 130.,
'force_symmorphic': True,
'nbnd': 20
},
'ELECTRONS': {
'mixing_mode': 'plain',
'mixing_beta': 0.7,
'conv_thr': 1.e-8,
'diago_thr_init': 5.0e-6,
'diago_full_acc': True
},
}
parameter_scf = Dict(dict=params_scf)
params_nscf = {
'CONTROL': {
'calculation': 'nscf',
'verbosity': 'high',
'wf_collect': True
},
'SYSTEM': {
'ecutwfc': 130.,
'force_symmorphic': True,
'nbnd': 500
},
'ELECTRONS': {
'mixing_mode': 'plain',
'mixing_beta': 0.6,
'conv_thr': 1.e-8,
'diagonalization': 'david',
'diago_thr_init': 5.0e-6,
'diago_full_acc': True
},
}
parameter_nscf = Dict(dict=params_nscf)
KpointsData = DataFactory('array.kpoints')
kpoints = KpointsData()
kpoints.set_kpoints_mesh([6,6,2])
alat = 2.4955987320 # Angstrom
the_cell = [[1.000000*alat, 0.000000, 0.000000],
[-0.500000*alat, 0.866025*alat, 0.000000],
[0.000000, 0.000000, 6.4436359260]]
atoms = Atoms('BNNB', [(1.2477994910, 0.7204172280, 0.0000000000),
(-0.0000001250, 1.4408346720, 0.0000000000),
(1.2477994910, 0.7204172280, 3.2218179630),
(-0.0000001250,1.4408346720, 3.2218179630)],
cell = [1,1,1])
atoms.set_cell(the_cell, scale_atoms=False)
atoms.set_pbc([True,True,True])
StructureData = DataFactory('structure')
structure = StructureData(ase=atoms)
params_gw = {
'HF_and_locXC': True,
'dipoles': True,
'ppa': True,
'gw0': True,
'em1d': True,
'Chimod': 'hartree',
#'EXXRLvcs': 40,
#'EXXRLvcs_units': 'Ry',
'BndsRnXp': [1, 10],
'NGsBlkXp': 2,
'NGsBlkXp_units': 'Ry',
'GbndRnge': [1, 10],
'DysSolver': "n",
'QPkrange': [[1, 1, 8, 9]],
'DIP_CPU': "1 1 1",
'DIP_ROLEs': "k c v",
'X_CPU': "1 1 1 1",
'X_ROLEs': "q k c v",
'SE_CPU': "1 1 1",
'SE_ROLEs': "q qp b",
}
params_gw = Dict(dict=params_gw)
builder = YamboConvergence.get_builder()
##################scf+nscf part of the builder
builder.ywfl.scf.pw.structure = structure
builder.ywfl.scf.pw.parameters = parameter_scf
builder.kpoints = kpoints
builder.ywfl.scf.pw.metadata.options.max_wallclock_seconds = \
options['max_wallclock_seconds']
builder.ywfl.scf.pw.metadata.options.resources = \
dict = options['resources']
if 'queue_name' in options:
builder.ywfl.scf.pw.metadata.options.queue_name = options['queue_name']
if 'qos' in options:
builder.ywfl.scf.pw.metadata.options.qos = options['qos']
if 'account' in options:
builder.ywfl.scf.pw.metadata.options.account = options['account']
builder.ywfl.scf.pw.metadata.options.custom_scheduler_commands = options['custom_scheduler_commands']
builder.ywfl.nscf.pw.structure = builder.ywfl.scf.pw.structure
builder.ywfl.nscf.pw.parameters = parameter_nscf
builder.ywfl.nscf.pw.metadata = builder.ywfl.scf.pw.metadata
builder.ywfl.scf.pw.code = load_node(options['pwcode_pk'])
builder.ywfl.nscf.pw.code = load_node(options['pwcode_pk'])
builder.ywfl.scf.pw.pseudos = validate_and_prepare_pseudos_inputs(
builder.ywfl.scf.pw.structure, pseudo_family = Str(options['pseudo_family'])
builder.ywfl.nscf.pw.pseudos = builder.ywfl.scf.pw.pseudos
##################yambo part of the builder
builder.ywfl.yres.yambo.metadata.options.max_wallclock_seconds = \
options['max_wallclock_seconds']
builder.ywfl.yres.yambo.metadata.options.resources = \
dict = options['resources']
if 'queue_name' in options:
builder.ywfl.yres.yambo.metadata.options.queue_name = options['queue_name']
if 'qos' in options:
builder.ywfl.yres.yambo.metadata.options.qos = options['qos']
if 'account' in options:
builder.ywfl.yres.yambo.metadata.options.account = options['account']
builder.ywfl.yres.yambo.parameters = params_gw
builder.ywfl.yres.yambo.precode_parameters = Dict(dict={})
builder.ywfl.yres.yambo.settings = Dict(dict={'INITIALISE': False, 'COPY_DBS': False})
builder.ywfl.yres.max_iterations = Int(5)
builder.ywfl.yres.yambo.preprocessing_code = load_node(options['yamboprecode_pk'])
builder.ywfl.yres.yambo.code = load_node(options['yambocode_pk'])
builder.parent_folder = load_node(options['parent_pk']).outputs.remote_folder
builder.precalc = builder.ywfl #for simplicity, to specify if PRE_CALC is True
builder.workflow_settings = Dict(dict={'type':'2D_space','what':'gap','where':[(1,8,1,9)],
'where_in_words':['Gamma'],'PRE_CALC':False})
#'what': 'single-levels','where':[(1,8),(1,9)]
para_space = [{'var':['BndsRnXp','GbndRnge'],
'space': [[[1,10],[1,10]], \
[[1,50],[1,75]], \
[[1,75],[1,50]]], \
'max_iterations': 1,},
{'var':['BndsRnXp','GbndRnge'],
'space': [[[1,75],[1,75]]],
'max_iterations': 1,}]
for i in range(len(para_space)):
print('{}-th variable will be {}'.format(i+1,para_space[i]['var']))
print('{}-th values will be {}'.format(i+1,para_space[i]['space']))
builder.parameters_space = List(list = para_space)
return builder
if __name__ == "__main__":
options = get_options()
builder = main(options)
running = submit(builder)
print("Submitted YamboConvergence; with pk=<{}>".format(running.pk))
'''
another example of space can be:
para_space = [{'var':['BndsRnXp','GbndRnge','NGsBlkXp'],
'space': [[[1, 50], [1, 50], 2], [[1, 50], [1, 50], 3]],'max_iterations': 1,},
{'var':['BndsRnXp','GbndRnge','NGsBlkXp'],
'space': [[[1, 55], [1, 55], 2], [[1, 55], [1, 55], 3]],'max_iterations': 1,},]
'''
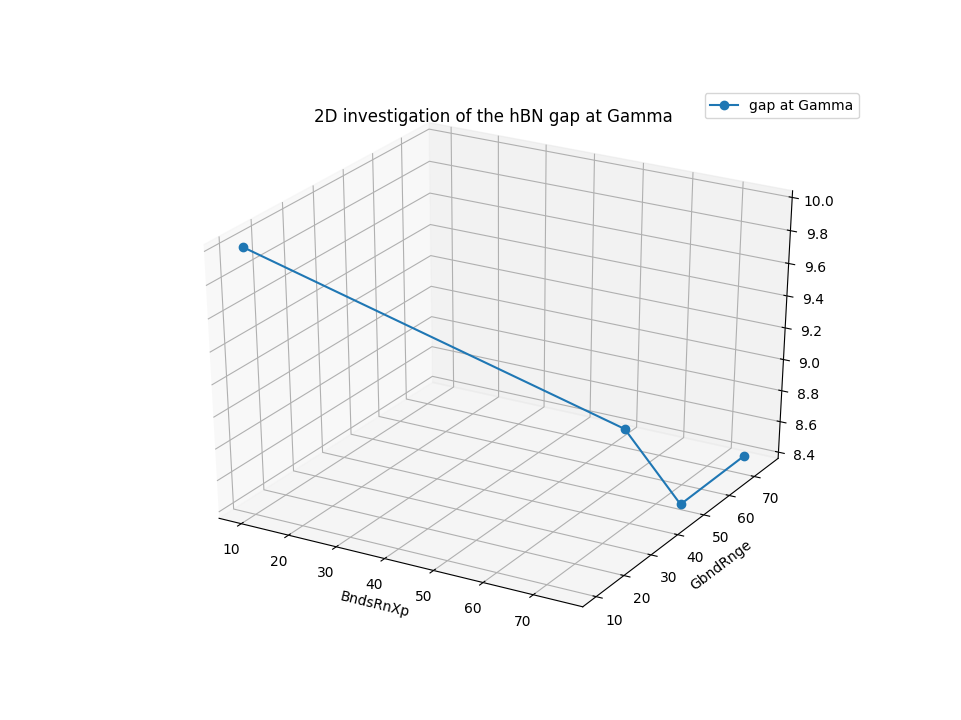
The type is called ‘2D_space’, but actually is possibile to change and arbitrary number of parameter at the time: the reason of the name is due to the fact that typically this investigation are done to observe the interdependence between bands and G-vectors cutoff. To see how to post process and plot results, we suggest to consult the dedicated section Plot results of YamboConvergence - fixed-path(‘2D_space’).